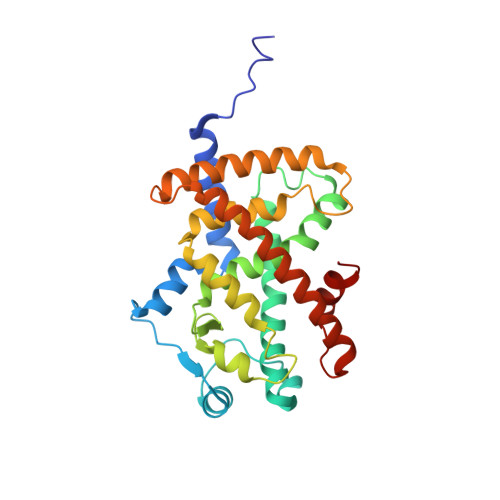
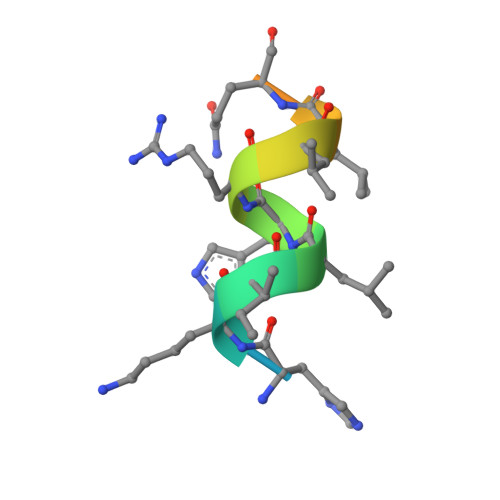
Structural basis for the inhibitory effects of a novel reversible covalent ligand on PPAR gamma phosphorylation.
Jang, J.Y., Kim, H., Kim, H.J., Suh, S.W., Park, S.B., Han, B.W.(2019) Sci Rep 9: 11168-11168
- PubMed: 31371757
- DOI: https://doi.org/10.1038/s41598-019-47672-w
- Primary Citation of Related Structures:
6IJR, 6IJS, 6JQ7 - PubMed Abstract:
Peroxisome proliferator-activated receptor γ (PPARγ) is a major therapeutic target for the treatment of type 2 diabetes. However, the use of PPARγ-targeted drugs, such as rosiglitazone and pioglitazone, is limited owing to serious side effects caused by classical agonism. Using a rational drug discovery approach, we recently developed SB1495, a novel reversible covalent inhibitor of the cyclin-dependent kinase 5 (Cdk5)-mediated phosphorylation of PPARγ at Ser245, a key factor in the insulin-sensitizing effect of PPARγ-targeted drugs. In this study, we report the crystal structures of PPARγ in complex with SB1495 and its enantiomeric analogue SB1494, which rarely exhibits inhibitory activity, to visualize the mechanistic basis for their distinct activities. SB1495 occupies the Arm3 region near the Ω loop of the PPARγ ligand-binding domain, whereas its enantiomeric analogue SB1494 binds to the Arm2 region. In addition, the piperazine moiety of SB1495 directly pushes the helix H2', resulting in the stabilization of the Ω loop just behind the helix H2'. Our results may contribute to the development of a new generation of antidiabetic drugs that selectively block PPARγ phosphorylation without classical agonism.
Organizational Affiliation:
Research Institute of Pharmaceutical Sciences, College of Pharmacy, Seoul National University, Seoul, 08826, Republic of Korea.